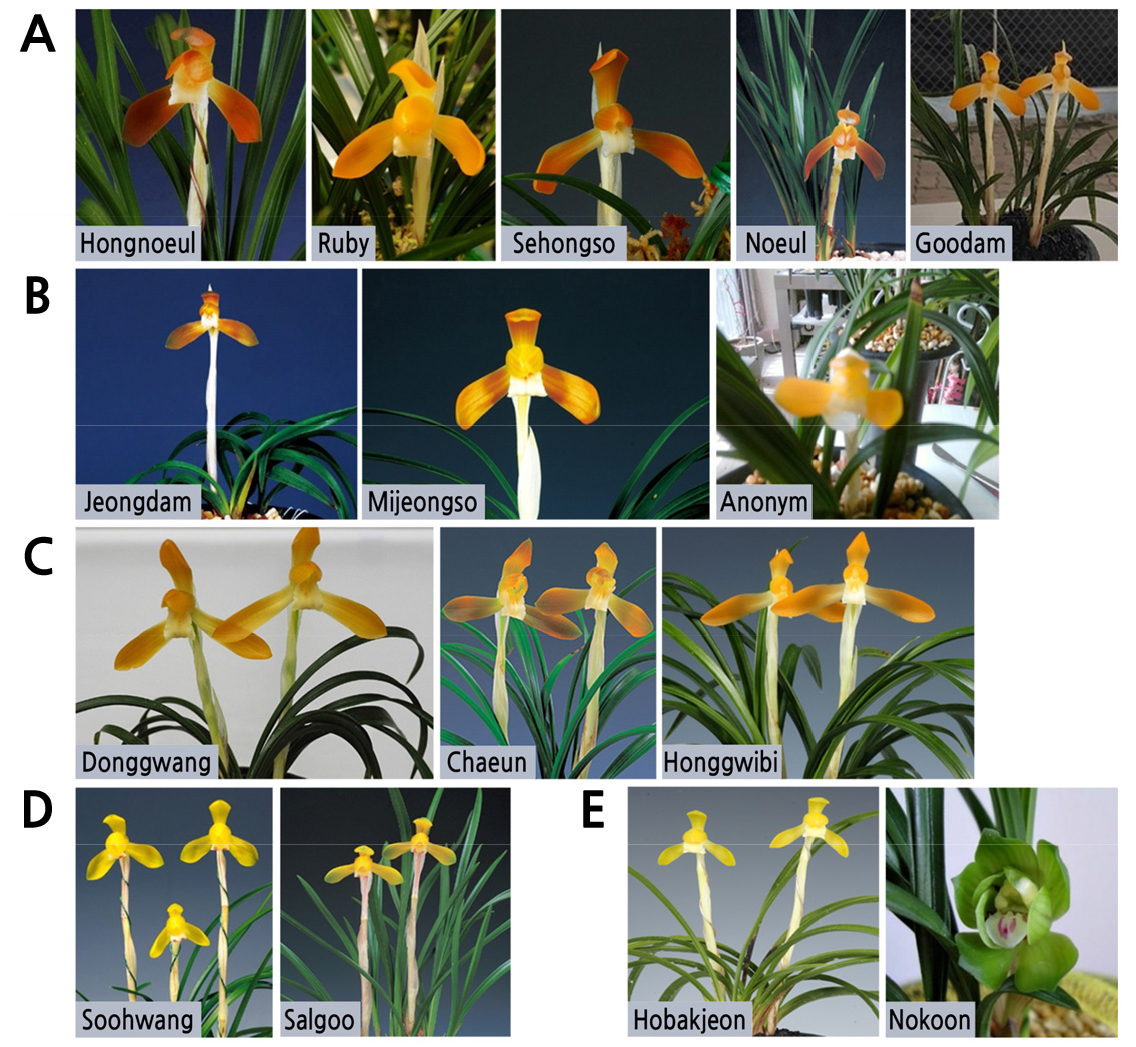

Y-chromosomal and mtDNA haplogroups were almost equally distributed between Western and Eastern Eurasian haplogroups. Inversely, various mtDNA lineages can be found on each site. Y-chromosome analysis revealed a burial pattern linked to paternal lineages, with men bearing closely related Y-haplotypes buried on the same sites. When related, the individuals were buried together, except for one adult woman, buried separately from her mother and young sister. Thirteen of the 28 individuals tested were linked by first-degree relationships. Familial relationships were assessed using the Likelihood Ratio (LR) method. Twenty-eight Scythian individuals from 5 archeological sites in the Tuva Republic (Russia) were analyzed using autosomal Short Tandem Repeats (STR), Y-STR and Y-SNP typing as well as whole mitochondrial (mtDNA) genome sequencing. In this work, we first aimed to address the question of the familial and social organization of some Scythian groups (Scytho-Siberians) by testing genetic kinship and, second, to add new elements on their origins through phylogeographical analyses.

Three hypotheses prevail regarding their origins that can be summarized as a “western origin”, an “eastern origin” and a “multi-regional origin”. However, their origins and the exact nature of their social organization remain debated. Scythians are known from written sources as horse-riding nomadic peoples who dominated the Eurasian steppe throughout the Iron Age.
Genodive genetic kinship mac os x#
genodive is available for computers running Mac OS X 10.7 or higher and can be downloaded freely from. In addition, genodive makes it possible to run several external programs ( lfmm, structure, instruct and vegan) directly from its own user interface, avoiding the need for data reformatting and use of the command line. A unique feature of genodive is that it can also open data sets with nongenetic variables, for example environmental data or geographical coordinates that can be included in the analysis. The different types of analyses offered by genodive include multiple statistics for estimating population differentiation ( φ ST, F ST, Fʹ ST, G ST, Gʹ ST, Gʹʹ ST, D est, R ST, ρ), analysis of molecular variance-based K-means clustering, Hardy–Weinberg equilibrium, hybrid index, population assignment, clone assignment, Mantel test, Spatial Autocorrelation, 23 ways of calculating genetic distances, and both principal components and principal coordinates analyses. One major feature of genodive is that it supports both diploid and polyploid data, up to octaploidy (2 n = 8 x) for some analyses, but up to hexadecaploidy (2 n = 16 x) for other analyses. Furthermore, genodive seamlessly supports 15 different file formats for importing or exporting data from or to other programs. genodive has an intuitive graphical user interface that allows direct manipulation of the data through transformation, imputation of missing data, and exclusion and inclusion of individuals, population and/or loci.

Genodive genetic kinship update#
This version presents a major update from the previous version and now offers a wide spectrum of different types of analyses. Genodive version 3.0 is a user-friendly program for the analysis of population genetic data.
